Webinar

LIMS Implementation: Avoiding Common Pitfalls and Fast Tracking Productivity
LIMS Implementation: Avoiding Common Pitfalls and Fast Tracking ProductivityBlog Post

How To Choose the Right ELN Vendor for Your Lab
How To Choose the Right ELN Vendor for Your LabWebinar

Why Sample Freezer Maps Matter—and How to Do It Right
Why Sample Freezer Maps Matter—and How to Do It RightBlog Post
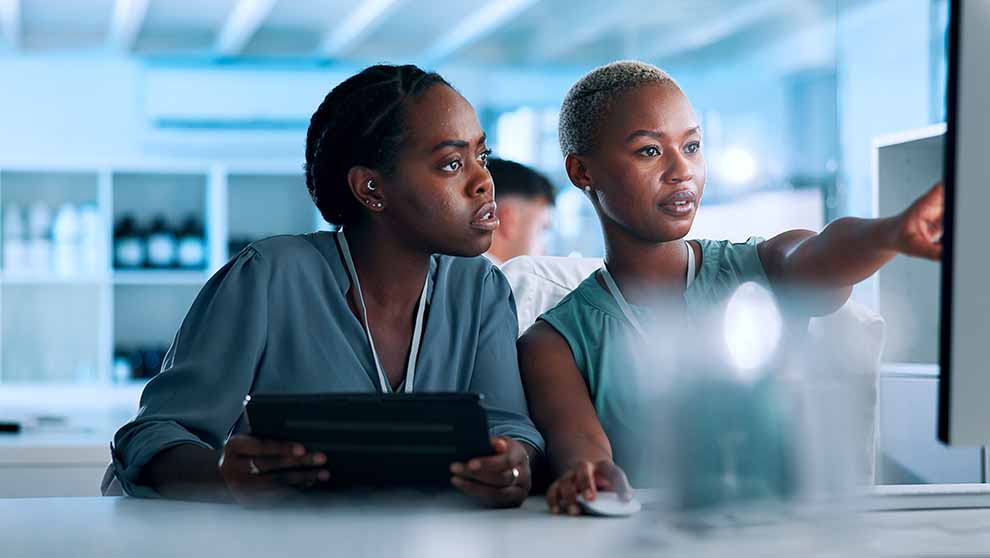

Why Lab Inventory Software Can Fail After Implementation (And How to Avoid It)
Why Lab Inventory Software Can Fail After Implementation (And How to Avoid It)Blog Post
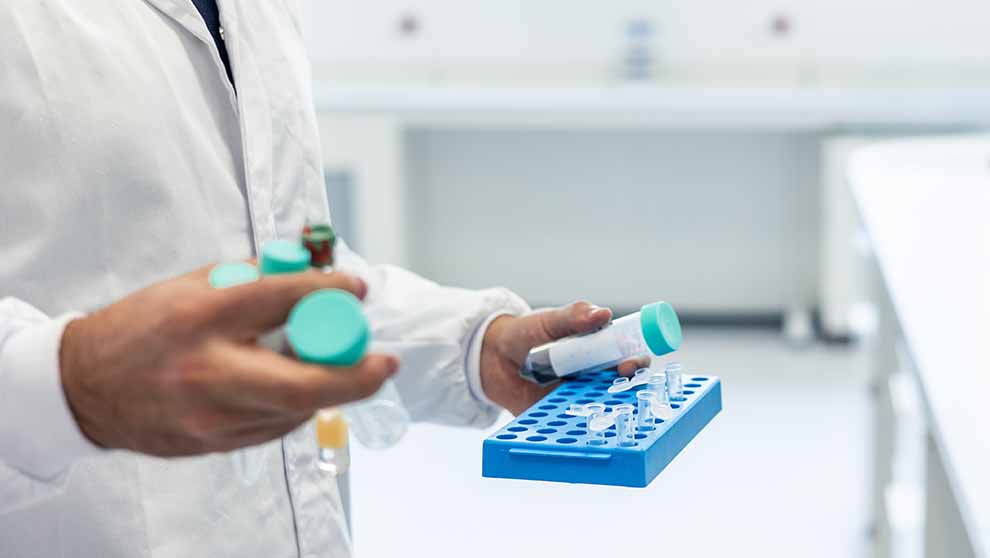

Get Synchronized: Biological Sample Storage for Multi-Site Labs and Biobanks
Get Synchronized: Biological Sample Storage for Multi-Site Labs and BiobanksBlog Post

Biologics Software: Tools for Emerging Biotechs
Biologics Software: Tools for Emerging BiotechsFeatured Content:
Ready for a demo?
Fill out the form and we will be in touch with you shortly to schedule your demo.